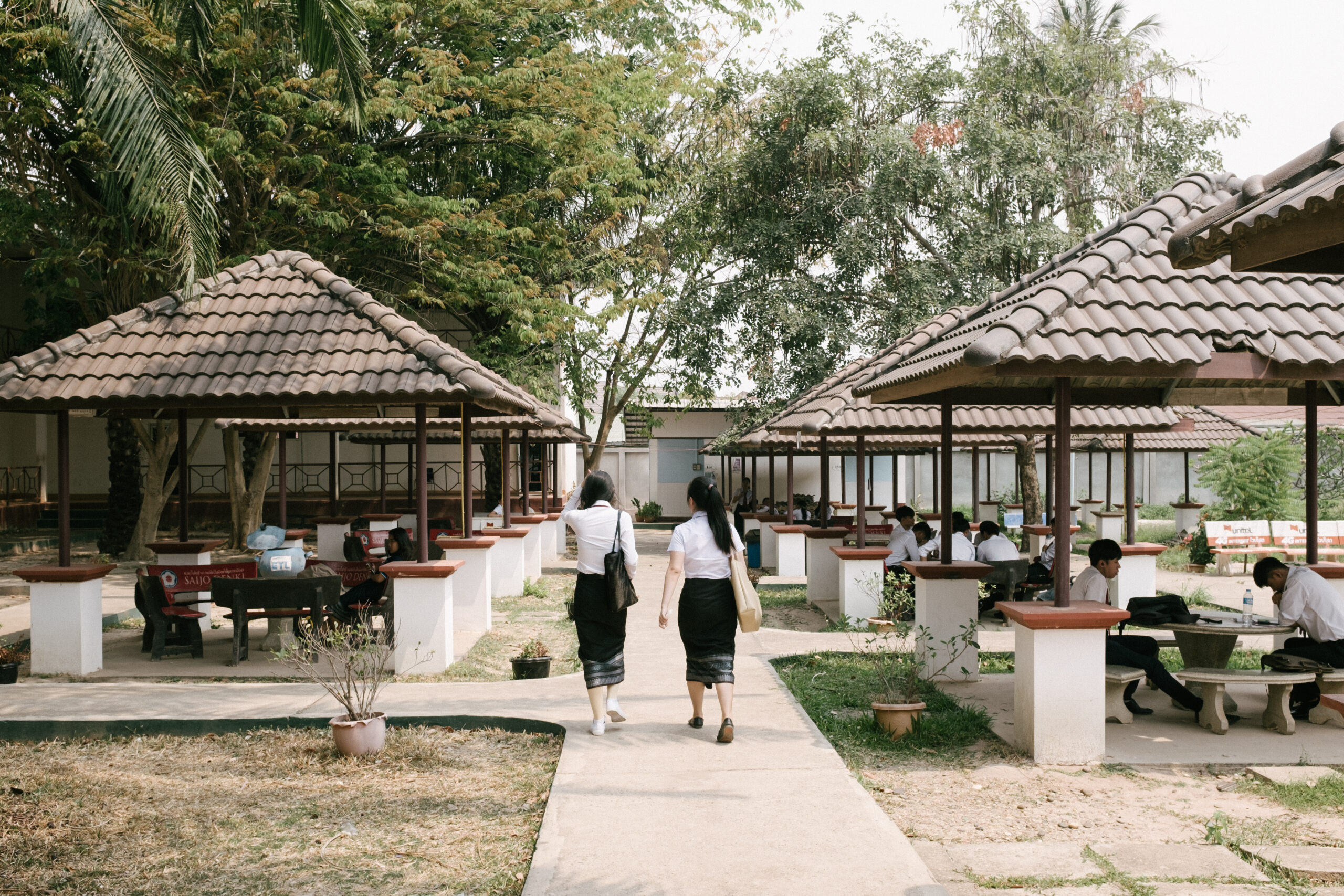
Aktualitäten
-

-
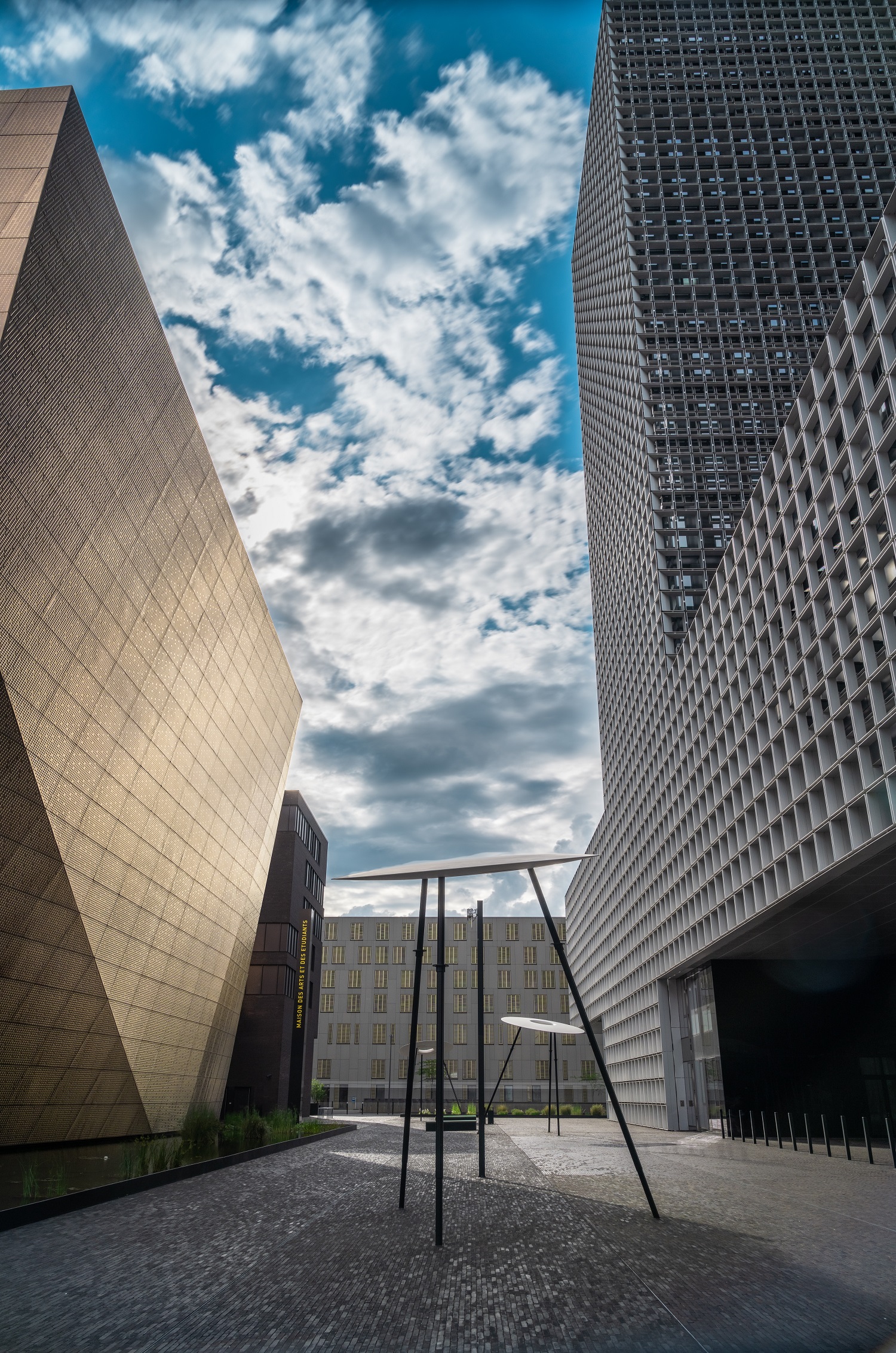

Uni.lu-Aktionsplan zur Förderung der Forschungsevaluierung jetzt verfügbar
Forschung, UniversitätMehr erfahren -

-
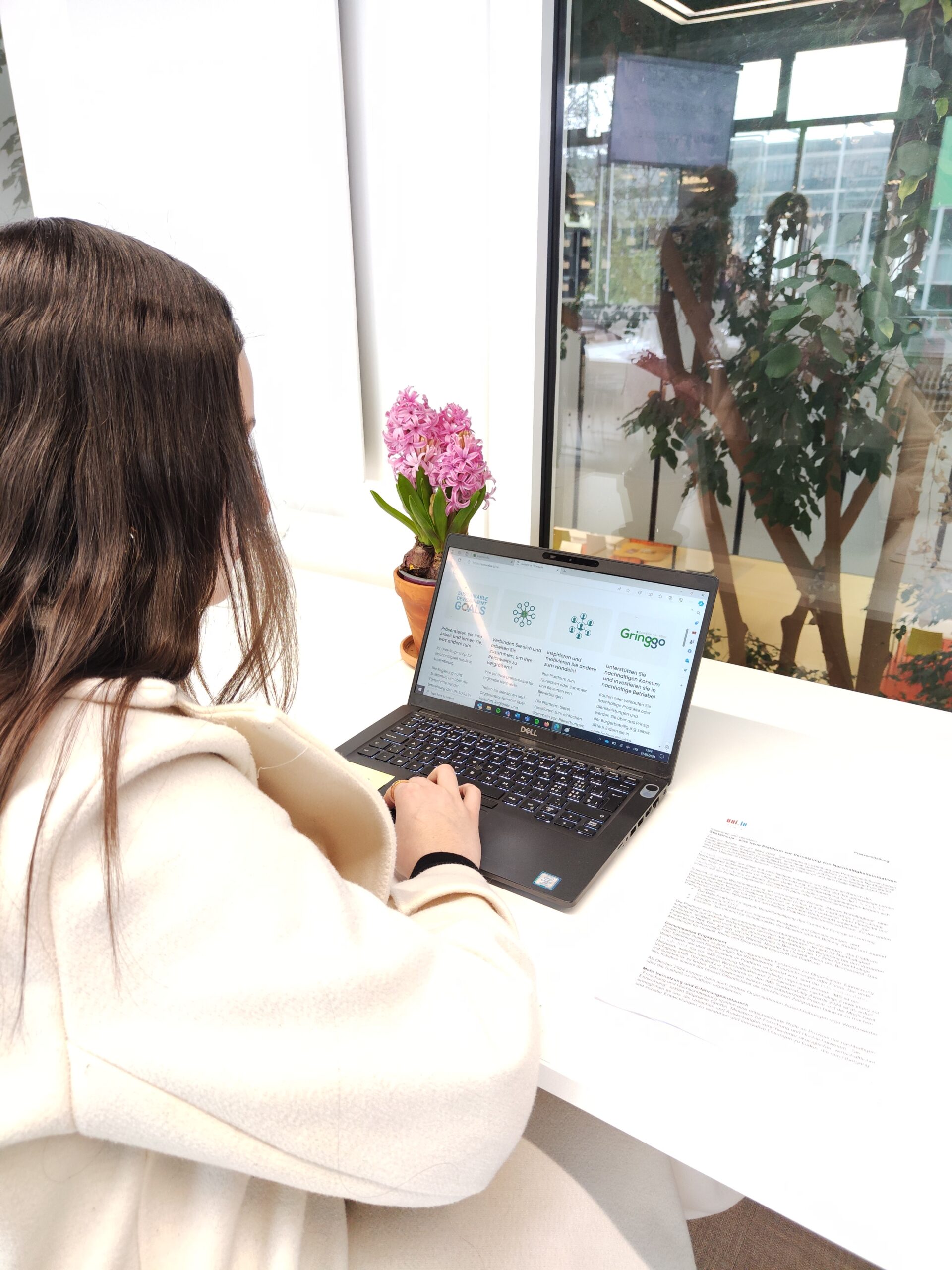

SustainLux – eine neue Plattform zur Vernetzung von Nachhaltigkeitsinitiativen
OutreachSozialwissenschaftenMehr erfahren -

-
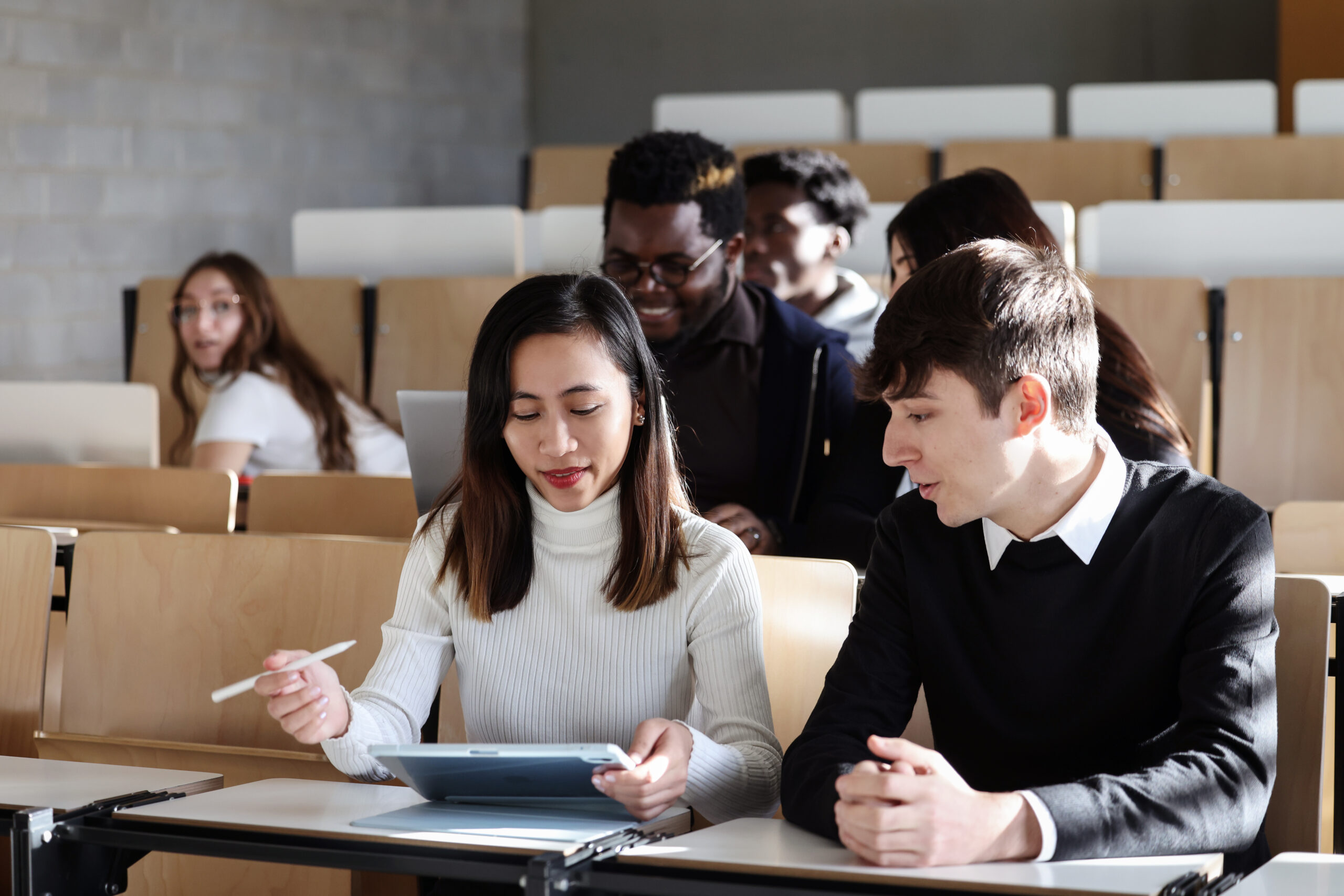

Neues im Studienangebot für den Studienbeginn 2024-2025
Studium, UniversitätGeisteswissenschaften, Ingenieurwissenschaften, Lebenswissenschaften & Medizin, RechtswissenschaftMehr erfahren
Aufruf zur Bewerbung
-

-

Appel à candidatures : Directeur fondateur /Directrice fondatrice du Centre Interdisciplinaire en Droit Européen
UniversitéMehr erfahren