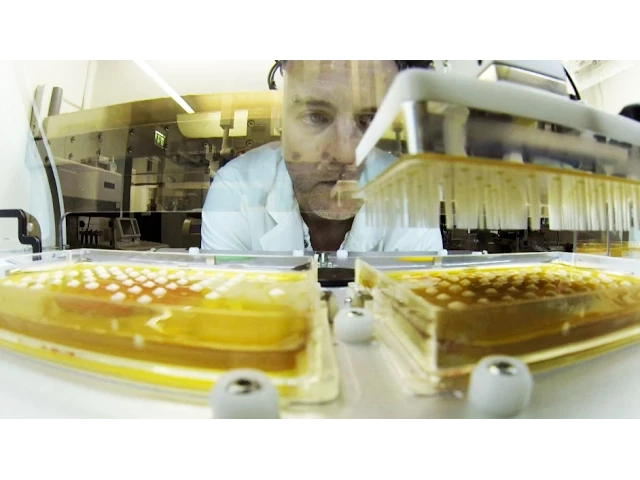
Einblicke in unsere Forschung
Unsere Forschung konzentriert sich auf neurodegenerative Erkrankungen wie Alzheimer oder Parkinson. Wann beginnen sie? Welche Rolle spielt die Genetik? Wie werden sie durch Lebensstil, Ernährung oder Umwelt beeinflusst? Wie wirken diese Faktoren zusammen? Wie lässt sich das Fortschreiten neurodegenerativer Erkrankungen verlangsamen oder gar verhindern? Wir suchen unermüdlich nach Antworten – zum Wohle der Patienten.
Das LCSB in Zahlen:
-
270Mitarbeiter
-
18Forschungsgruppen
Vorträge
 Donnerstag 16 MaiKostenfrei, Präsenzveranstaltungen, Vorträge
Donnerstag 16 MaiKostenfrei, Präsenzveranstaltungen, VorträgeAlzheimer-Krankheit: Wie sie verhindert werden kann!
Mehr erfahren Dienstag 23 AprilKostenfrei, Präsenzveranstaltungen, Vorträge
Dienstag 23 AprilKostenfrei, Präsenzveranstaltungen, VorträgeForschung für Parkinson-Prävention und Präzisionsmedizin
Mehr erfahren

